
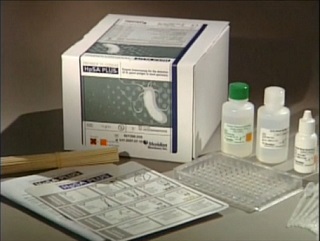
Helicobacter pylori antigen test was performed using Premier Platinum HpSA PLUS enzyme
immunoassay (EIA). The kit used was purchased from Meridian Bioscience Inc. The
Premier Platinum HpSA PLUS enzyme immunoassay (EIA) kit is an in vitro qualitative procedure for the
detection of Helicobacter pylori antigens in human stool. Test results
are intended to aid in the diagnosis of H. pylori infection and to
monitor response during and post-therapy in patients. Accepted medical practice
recommends that testing by any current method, to confirm eradication, be done
at least four weeks following completion of therapy.
Helicobacter pylori which
is previously known as Campylobacter pylori, is a gram negative, microaerophilic bacterium usually found in the stomach.
It was identified in 1982 by Australian scientists Barry
Marshall and Robin Warren,
who found that it was present in a person with chronic gastritis and gastric
ulcers, conditions not previously believed to have a microbial cause. It is also linked to the
development of duodenal ulcers and stomach
cancer. However, over 80% of individuals infected with the bacterium
are asymptomatic,
and it may play an important role in the natural stomach ecology. According to research,
more than 50% of the world's population harbour H. pylori in their upper gastrointestinal tract. Infection is more
prevalent in developing countries, and incidence is decreasing in Western
countries. H. pylori's helical shape and its flagella is used
to penetrate the mucoid lining of the stomach to reach the epithelial cells underneath,
where the pH is more neutral. They also neutralise the acid in its
environment by producing large amounts of urease, which breaks down the urea present in the stomach to carbon dioxide and ammonia. The ammonia, which is basic, then neutralizes stomach acid.
This ammonia produced to regulate pH is toxic to epithelial cells, as well as
other biochemicals produced by H. pylori such as proteases, vacuolating cytotoxin A (VacA) and certain phospholipases. Cytotoxin associated
gene CagA can
also cause inflammation and is potentially a carcinogen.
Symptoms
Most people infected with H. pylori are asymptomatic but in acute
infection may appear as an acute gastritis with abdominal pain or nausea. If the
infection develops into chronic gastritis, the symptoms may include nausea,
belching, stomach pains, bloating, black stool
and sometimes vomiting.
Individuals infected with H. pylori have a 10 - 20% lifetime risk of
developing peptic ulcers and a 1 - 2% risk of acquiring stomach
cancer. Inflammation
of the pyloric
antrum is more likely
to lead to duodenal ulcers, while inflammation of the corpus (body of the stomach) is more likely
to lead to gastric ulcers and gastric carcinoma.
However, H. pylori possibly
plays a role only in the first stage that leads to common chronic inflammation,
but not in further stages leading to carcinogenesis. A meta-analysis conducted in 2009
concluded the eradication of H.
pylori reduces gastric cancer
risk in previously infected individuals, suggesting the continued presence of H. pylori constitutes a relative risk factor of 65% for gastric cancers and in
terms of absolute risk,
the increase was from 1.1 - 1.7%. H. pylori have been associated with colorectal
polyps and colorectal
cancer and may also be associated with eye disease
SPECIMEN
COLLECTION AND PREPARATION
The faeces
sample should be received in an airtight transport container and stored at 2-8oC
until tested. The specimen should be tested as soon as possible, but may be
held up to 72 hours at 2-8oC prior to testing. If testing cannot be
performed within this time frame, specimens should be frozen immediately upon
receipt and stored frozen (-20oC to –80oC) until tested.
Specimens may be frozen and thawed twice.
Stool
in transport media, swabs, or preservatives are inappropriate for testing.
SPECIMEN
PREPARATION
1.
Using a pipetting device, add 500μL of Sample Diluent to a clean test tube.
2.
Mix stool as thoroughly as possible prior to pipetting.
a.
Liquid or semi-solid stools - Using the supplied transfer pipette, add 100μL (second
mark from the tip of the pipette) of stool into Sample Diluent. Using same
pipette, gently withdraw and expel the stool suspension several times, then
vortex 15 seconds. Save the transfer pipette in the sample for later use.
b.
Formed/Solid stools - Using a wooden applicator stick, transfer a small portion
(5-6 mm diameter) of thoroughly mixed stool into Sample Diluent. Emulsify stool
using the wooden applicator stick, then vortex 15 seconds.
3.
Stool specimens may be centrifuged after dilution. Centrifuge at approximately
2750
x G for five minutes or until solid matter separates from liquid. Proceed with the
assay after recovering supernate.
TEST
PROCEDURE
1.
After the pouch has reached temperature, break off the required number of
microwells
(1 well for each specimen, plus 1 positive and 1 negative control well per
batch).
Place the microwells in the microwell strip holder and record the location of
all
wells. Unused microwells must be resealed in the pouch immediately.
2.
Using the specimen transfer pipette, add 100μL of diluted stool (second mark
from the tip of the pipette) to the appropriate well. (Place the pipette tip
halfway into well and let the sample slowly run down side of well.)
3.
Add 2 free falling drops of Positive Control and 100μL of Sample Diluent /Negative
Control to the appropriate wells.
4.
Add 1 free falling drop (approximately 50μL) of Enzyme Conjugate to each well.
Firmly
shake/swirl the plate for 30 seconds.
5.
Cut plate sealer to size and press firmly onto top of microwells to seal.
Incubate the plate for 1 hour at 19-27oC
6.
Carefully remove the plate sealer and wash wells:
a.
Manual method:
i
Dump plate contents firmly into a biohazard receptacle.
ii
Bang the inverted plate on a clean stack of paper towels.
iii
Fill all wells with 1X Wash Buffer I, directing stream of buffer to the sides of
the wells to avoid foaming.
iv
Repeat wash cycle (dump, bang on fresh towels, fill) 4 times for a total of 5
wash cycles. After the last fill, dump and bang plates on fresh towels hard
enough to remove as much excess wash buffer as possible, but do not allow wells
to completely dry at any time.
b.
Semiautomated method using validated equipment
i
Aspirate the contents of the well.
ii
Fill the wells to the top (approx. 300-350μL/well) with 1X Wash Buffer I then
aspirate. The washer manifold should be adjusted to ensure no foaming occurs
during the filling of the wells and that the wells are thoroughly aspirated
after each wash.
iii
Repeat step ii a minimum of 4 more times. Following the last wash, test wells should
be thoroughly aspirated to remove as much moisture as possible.
7.
Clean the underside of all wells with a lint-free tissue.
8.
Add 2 free falling drops (approx. 100μL) of Premier Substrate Solution I to
each well. Firmly shake/swirl the plate for 30 seconds. Incubate for 10 minutes
at 19-27 oC.
9.
Add 2 free-falling drops (approx. 100μL) of Premier Stop Solution I to each
well.
Firmly
shake/swirl the plate for 30 seconds.
Initial
colour of positive reaction is blue, which changes to yellow upon addition
of
Premier Stop Solution I.
10.
Inspect and record reactions. Test results can be read visually or using a
spectrophotometric
reader.
a.
Visual Determination - Read within 15 minutes after adding Premier Stop
Solution
I.
b.
Spectrophotometric Determination - Zero EIA reader on air. Wipe underside
of
wells with a lint-free tissue. Read absorbance at 450 nm or 450/630 nm
within
15 minutes of adding Premier Stop Solution I.
INTERPRETATION
OF RESULTS
The
following interpretations apply to both initial diagnosis and monitoring of
anti-H. pylori therapy.
Visual
Reading
Negative
= colorless to faint yellow
Positive
= definite yellow color
To
be called positive, a faint yellow color must be confirmed by a spectrophotometric
reading. If a spectrophotometer is not available, the cut-off must be
determined by an alternative method.
Spectrophotometric
Single Wavelength (450 nm)
Negative:
< 0.140
Positive:
≥ 0.140
Negative
Control: < 0.140
Positive Control: ≥ 0.640
No comments:
Post a Comment